
Pathogenic variants ranked in first positions

Classification accuracy up to 98%

VUS reduction up to 60%

Intuitive and Fast

Secure and CE IVD marked
Winner of the NIH-funded CAGI6 Challenge, eVai platform automates ACMG guidelines and prioritizes variants to highlight candidate diagnoses.
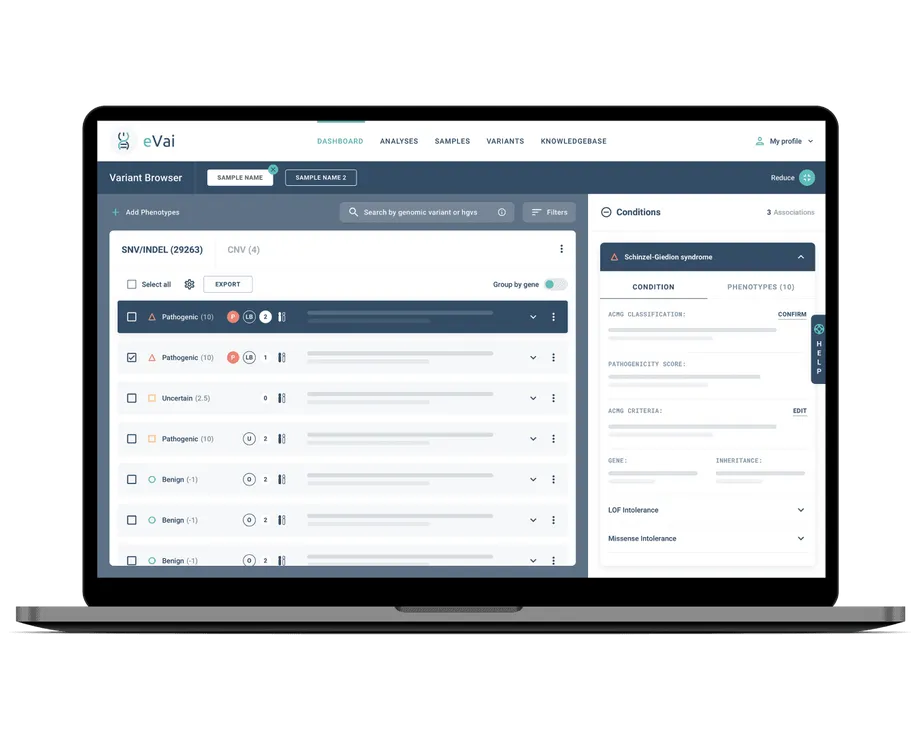


Pathogenic variants ranked in first positions

Classification accuracy up to 98%

VUS reduction up to 60%

Intuitive and Fast

Secure and CE IVD marked
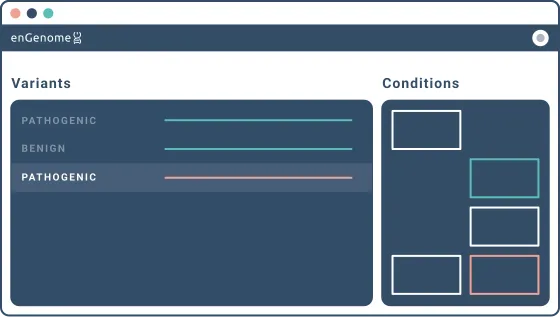
eVai automatically pre-classifies SNV, Indels and CNVs and ranks them according to a hypothesis-free approach. The classification is disease-specific and reports both the activated criteria and the supporting evidence.
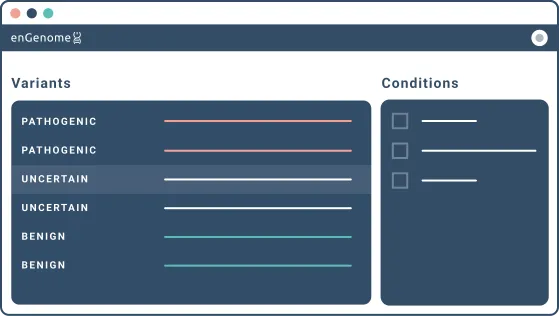
eVai leverages a proprietary AI technology that exploits variant pathogenicity together with patient phenotypes and family information to suggest the most likely genetic diagnosis for the patient.

Our Suggested Diagnosis can predict digenic and oligogenic combinations based on a rigorously evaluated and benchmarked model. Users can explore candidate pairs and the supporting literature if available.

Variant-disease associations can be submitted to a private lab repository that can contribute to significantly improve classification and prioritization performances over time. Moreover, the lab repository can be used to store info about variant artifacts.
Analyze trio, quartet or up to 7 family members: for each variant, eVai will automatically infer the possible inheritance patterns according to a fully penetrant disease. Users can select the variants that fit certain inheritance patterns and quickly compare genotypes across the members.
eVai enables users to create virtual gene panels and custom variant filters by leveraging on multiple omics resources such as Human Phenotype Ontology.
Make your interpretation process intuitive, faster and more accurate.
VarChat is the first open platform to use Generative AI for searching genomic variants in literature and synthesizing information in a coherent and concise way.


